Solvation Theory
Mixing molecules and water, or solvating the molecules, changes the characteristics of the water (the solvent). We can use this information to predict the physical properties of and interactions between the molecules (the solute).
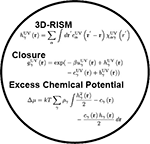
Predicting the distribution of the liquid and changes in thermodynamics can be done with different computational models, such as explicitly representing every atom of the liquid or treating it as a continuum dielectric. However, these approaches each have their own drawbacks: computation time, numerical noise, lack of physical detail or accuracy. To mitigate these drawbacks, our work focuses on the development and application of the 3D reference interaction site model (3D-RISM) theory of molecular solvation. 3D-RISM uses statistical physics to predict the molecular distribution of explicit, all-atom solvent models around solute molecules and calculate the solvation thermodynamics from these distributions. With 3D-RISM, we can, for example, predict the ion distributions around ions, water distribution around proteins, or solvation free energy of small molecules.
Since solvation is a change in the liquid from its bulk state, we also need to model the bulk liquid. RISM theory can be used to study polar liquids (like water), non-polar liquids (like cyclohexane), or mixtures (like ions in water). We are also working to improve RISM theory by proposing new approaches to fundamental approximations in the theory .